In my previous article, the R scripts were fairly simple. The more complex the R script is, the more specific your R computation will be. As you progress in your use of R, you will be adding references to new R packages. At some point, you will start using particular R packages that are designed for specific purposes. These could be for graphs, for connections to different data sources, for specific mathematical or statistical computations, or many others.
Essentially, an R package is a container of functions that serve a particular purpose, with the binary source code (usually C++), documentation, and sample data. A package is a collection of these files that reside in a library folder. If you navigate to your R library folder, you will see all the packages installed for your R engine. A package might also be called a binary package or a tarball, depending on the operating system.
A package is not equivalent to a library, nor should be mistaken for one. In the R language, when installing a package, the command "install.packages" is used. The word, library(), is a function that loads functions in particular packages into your R environment. Deriving from the Windows OS, shared objects are called DLLs (dynamic-link library), hence the word library is used and refers to common and shared objects. So, to load a package into your R environment, the function library() is used, with the name of package specified in brackets.
Referring to an R library in an R Script is super easy; simply add the library or use the require() method. But in the system the library must be, in the first place, installed. Following on from my previous article, Introduction to R Services, I will extend the example with sp_execute_external_script and we will also call new libraries.
Libraries are installed by installing packages available in common repositories, such as CRAN, Biocondutor, Github and many others. In the R language, a library is installed by invoking this command:
install.packages("name_of_the_package")Since installation of such packages is not possible by running an R script with external stored procedure, we will explore the correct way to do this. This code will return an error
--Install Package using sp_execute_external_script
EXECUTE sp_execute_external_script
@language = N'R'
,@script=N'install.packages("AUC")'This T-SQL code return an error, saying that this package is not available for my R version. But we will later see how to install same package.
So if we extend the original stored procedure to:
EXECUTE sp_execute_external_script
@language = N'R'
,@script = N' library(Hmisc)
u <- unlist(rcorr(Customers_by_invoices$InvoiceV, Customers_by_invoices$CustCat, type="spearman"))
statistical_significance <-as.character(u[10])
OutputDataSet <- data.frame(statistical_significance)'
,@input_data_1 = N'SELECT
SUM(il.Quantity) AS InvoiceQ
,SUM(il.ExtendedPrice) AS InvoiceV
,c.CustomerID AS Customer
,c.CustomerCategoryID AS CustCat
FROM sales.InvoiceLines AS il
INNER JOIN sales.Invoices AS i
ON il.InvoiceID = i.InvoiceID
INNER JOIN sales.Customers AS c
ON c.CustomerID = i.CustomerID
GROUP BY
c.CustomerID
,c.CustomerCategoryID'
,@input_data_1_name = N'Customers_by_invoices'
WITH RESULT SETS (( statistical_significance FLOAT(20) ));
GOWe will be able to calculate statistical significance for the correlation between two variables. Point given here is, that we are referring to R function “library(Hmisc)” with package name. See the detailed part of the script below.
-- part of R script with reference to call method library
,@script = N' library(Hmisc)
u <- unlist(rcorr(Customers_by_invoices$InvoiceV, Customers_by_invoices$CustCat, type="spearman"))
statistical_significance <-as.character(u[10])
OutputDataSet <- data.frame(statistical_significance)'When referring a library, we need to have package already preinstalled, otherwise, you will receive an error, saying that package “Hmisc” that we are referring to, does not exists and R script cannot be executed.
General information on packages
Before taking a look at different approaches on how to install missing packages, let's take a look where packages are saved. Packages are always saved in folder library, but depending on your version of R (Open, Client, Server), paths will be different.
For Client or Server version, you will find your libraries on your main drive. For Client version, the default path is: C:\Program Files\Microsoft\R Client\R_SERVER\library. You can see the folder contents in the image below.

And for R Server version, you will find libraries on path of your default SQL Server instance: C:\Program Files\Microsoft SQL Server\MSSQL13.MSSQLSERVER\R_SERVICES\library. Here are the contents from a server installation.
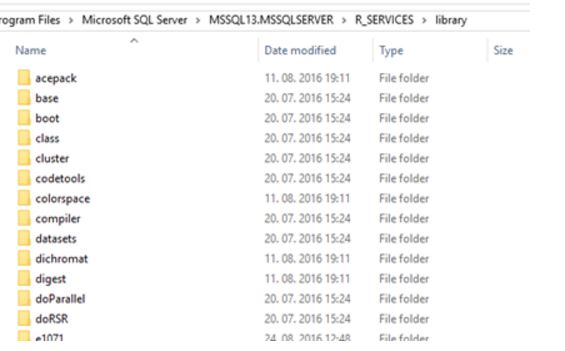
Subfolders represent the name of the installed and available packages. To make sure and find the default path to your packages, you can execute following code:
-- Path to libraries on your computer/server
EXECUTE sp_execute_external_script
@language = N'R'
,@script = N'OutputDataSet <- data.frame(.libPaths());'
WITH RESULT SETS (([DefaultLibraryName] VARCHAR(MAX) NOT NULL));
GOand in my case, this is the default path for R packages in R Server edition.
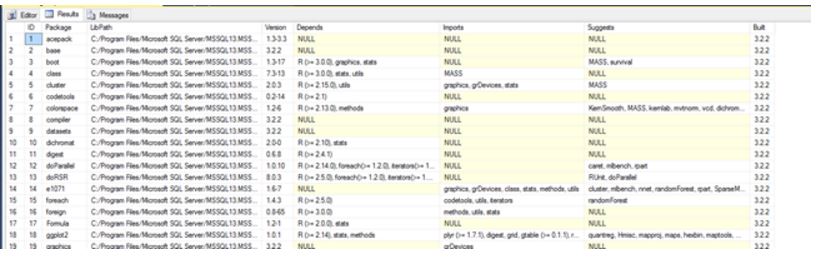
Much more information can be retrieved by using R function installed.packages(). In this example, we are extract much more information on packages and inserting the information into SQL Server table.
-- You can create a table for libraries and populate all the necessary information
CREATE TABLE dbo.Libraries
(
ID INT IDENTITY NOT NULL CONSTRAINT PK_RLibraries PRIMARY KEY CLUSTERED
,Package NVARCHAR(50)
,LibPath NVARCHAR(200)
,[Version] NVARCHAR(20)
,Depends NVARCHAR(200)
,Imports NVARCHAR(200)
,Suggests NVARCHAR(200)
,Built NVARCHAR(20)
)
INSERT INTO dbo.Libraries
EXECUTE sp_execute_external_script
@language = N'R'
,@script=N'x <- data.frame(installed.packages())
OutputDataSet <- x[,c(1:3,5,6,8,16)]'
SELECT * FROM dbo.Libraries
DROP TABLE dbo.LibrariesBy querying this table, you get information on library dependencies, versions, imports and builds in one execution of sp_execute_external_script.
In the following four ways (and I believe this number is not final), I will show, how you can install new R packages. The first two ways are proposed by Microsoft (as described on MSDN), but the last two I have found very useful and simple, especially when deploying or sharing T-SQL or R code. Since the last two ways are not the official ways (proposed by Microsoft) for installing packages, please take some caution if you use them.
Using R Tools for Visual Studio
This is recommended by Microsoft. The user should have installed RTVS (R Tools for Visual Studio) on client in order to install packages. In RTVS user can run couple of settings in order so see the default paths.
sessionInfo() .libPaths()
This also returns the paths to the library folders on my machine - for R Server (by default C:/Program Files/Microsoft SQL Server/MSSQL13.MSSQLSERVER/R_SERVICES/library) and for user (C:/Users/SI01017988/Documents/R/win-library/3.2) that is currently using RTVS.

By checking Options,
then the R Tools..Advanced (as shown below):
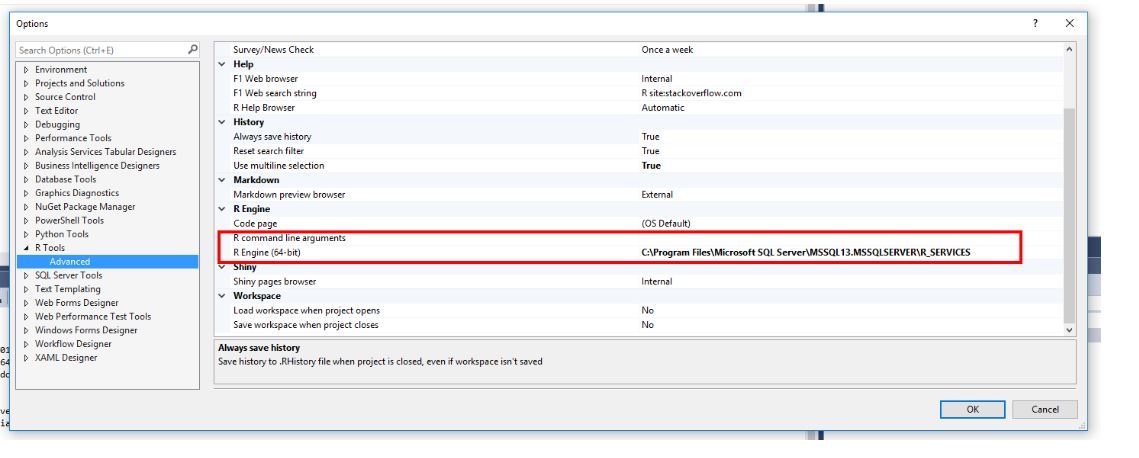
you will see that the R engine has the path defined. This path is the root path for installing R packages. R Packages are installed under the subfolder, Library (C:\Program Files\Microsoft SQL Server\MSSQL13.MSSQLSERVER\R_SERVICES\Library).
So by running install.packages("AUC") I can see that the server path was ignored and library was installed into users folder.
By using the library() function, I will check the presence of this package.
EXECUTE sp_execute_external_script
@language = N'R'
,@script = N'library("AUC")
OutputDataSet <- data.frame(.libPaths());'After running this, I received an error, stating "there is no package called 'AUC'", as if the installation I did just did not happen.
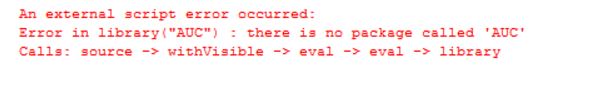
To install package in R Server folder you either need to have administration access or have an Administrator do it for you. Microsoft has discussed this issue on MSDN. Running Visual Studio and RTVS as an administrator does not change the result. You can see this in the following BOL article: Problems with Packages Installed in User Libraries. To avoid this problem, you need to declare the path to the library folder of R Server, where you want package to be installed.
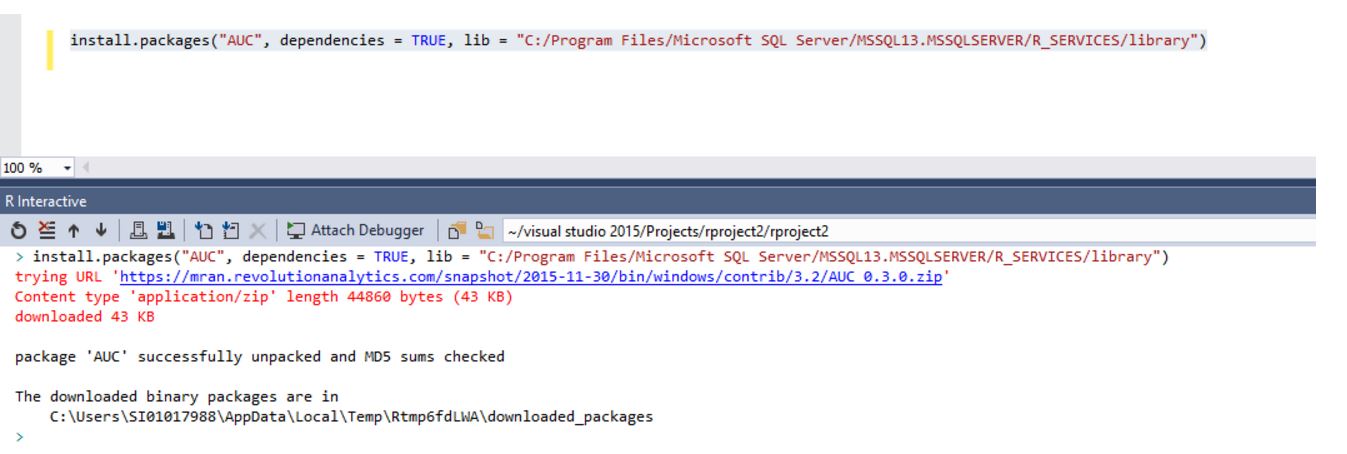
install.packages("AUC", dependencies = TRUE, lib = "C:/Program Files/Microsoft SQL Server/MSSQL13.MSSQLSERVER/R_SERVICES/library")And the package will be installed automatically R Server repository folder.
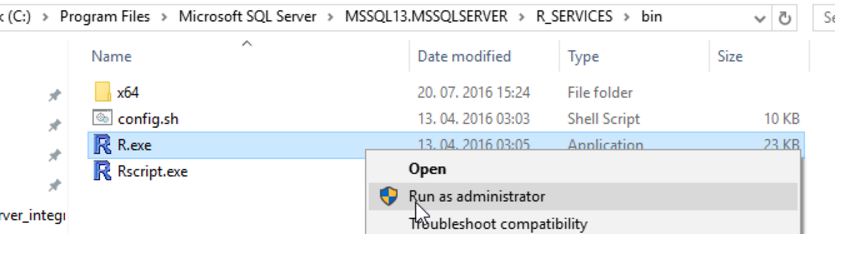
Using R.EXE in CMD
When executing R.EXE as administrator in the binary folder of R Server (C:\Program Files\Microsoft SQL Server\MSSQL13.MSSQLSERVER\R_SERVICES\bin), you will be prompted with a command window.
From there, simply execute:
install.packages("AUC")Using XP_CMDSHELL
This method is by far the fastest and very useful when sharing T-SQL code. By enabling xp_cmdshell in configurations, and using this stored procedurem, you can install any missing package. Using command code with switch -e, you can easily install the library.
R cmd -e install.packages("AUC")
The following code will install the missing package on R Server.
USE WideWorldImporters;
GO
-- enable xp_cmdshell
EXECUTE SP_CONFIGURE 'xp_cmdshell','1';
GO
RECONFIGURE;
GO
EXEC xp_cmdshell '"C:\Program Files\Microsoft SQL Server\MSSQL13.MSSQLSERVER\R_SERVICES\bin\R.EXE" cmd -e install.packages(''AUC'')';
GOThe results from the command are displayed in SSMS, as shown here.

It might not be the proposed way, but when completing T-SQL and R code it is just a nice way with few annoyances or switching between programs. I would still suggest the official way Microsoft suggests.
Using CMD and the -e switch, the deployment and computation of R code can be executed and simplified.
Copying Files
Copying files may sound a bit strange at the first glance but for majority of packages, copying the complete library folder to destination folder will also do the job. Image being a system administrator responsible for configuring a new R environment or sandbox. In this scenario you don't want to run hundreds of lines of code to install all the needed packages, but you would just copy/paste the folders from some image or backup. This would be 100x faster and more convenient. Also, the packages would be already tested with an R engine.
Using a simple copy / paste, drag and drop, I copied and "installed" the package, acepack, in my R Client environment (copied from R Server).
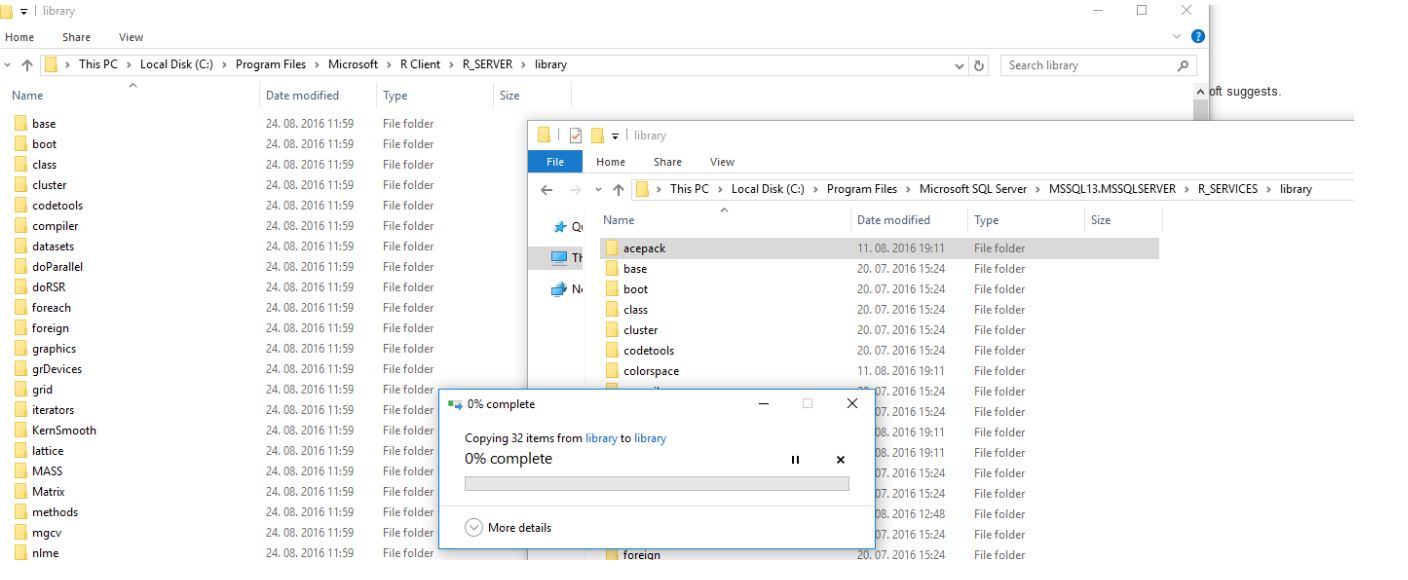
Conclusion
The installation of packages and referring to them should not be overlooked and must not slow down your daily process or using R script. To avoid any additional annoyances, when calling a specific package, make sure that you have in your R environment all the necessary packages. To have 1000 packages installed in an R environment is not unusual for a data scientist. In a production system, I would recommend to always have only those libraries that are needed for the execution of a particular script or job. And keep in mind, that compatibility of the R engine and packages can change from version to version. To avoid any unexpected event, have them tested in your test environment.
Author: Tomaz Kastrun (tomaz.kastrun@gmail.com)
Twitter: @tomaz_tsql


